
APA Style
Koki Yoshimoto, Ririka Okaichi, Kei Iida, Shiho Terada, Ken-ichiro Kamei. (2025). High-Purity Production of Endothelial Cells from Human Pluripotent Stem Cells. Cell Therapy & Engineering Connect, 1 (Article ID: 0006). https://doi.org/10.69709/CTEC.2025.154455MLA Style
Koki Yoshimoto, Ririka Okaichi, Kei Iida, Shiho Terada, Ken-ichiro Kamei. "High-Purity Production of Endothelial Cells from Human Pluripotent Stem Cells". Cell Therapy & Engineering Connect, vol. 1, 2025, Article ID: 0006, https://doi.org/10.69709/CTEC.2025.154455.Chicago Style
Koki Yoshimoto, Ririka Okaichi, Kei Iida, Shiho Terada, Ken-ichiro Kamei. 2025. "High-Purity Production of Endothelial Cells from Human Pluripotent Stem Cells." Cell Therapy & Engineering Connect 1 (2025): 0006. https://doi.org/10.69709/CTEC.2025.154455.
 ACCESS
Research Article
ACCESS
Research Article
Volume 1, Article ID: 2025.0006

Koki Yoshimoto
yoshimoto.koki.5a@kyoto-u.ac.jp

Ririka Okaichi
2210330032b@kindai.ac.jp

Kei Iida
kiida@life.kindai.ac.jp

Shiho Terada
sterada@icems.kyoto-u.ac.jp

Ken-ichiro Kamei
kk4801@nyu.edu
1 Institute for Integrated Cell-Material Sciences, Kyoto University Institute for Advanced Study, Kyoto 606-8501, Japan
2 Department of Life Science, Faculty of Science and Engineering, Kindai University, Higashiosaka 577-8502, Japan
3 Center for Cancer Immunotherapy and Immunobiology, Kyoto University Graduate School of Medicine, Kyoto 606-8501, Japan
4 Program of Biology, Division of Science, New York University Abu Dhabi, Abu Dhabi P.O. Box 129188, United Arab Emirates
5 Program of Bioengineering, Division of Engineering, New York University Abu Dhabi, Abu Dhabi P.O. Box 129188, United Arab Emirates
6 Department of Biomedical Engineering, Tandon School of Engineering, New York University, Brooklyn, NY 11201, USA
7 Department of Biology, Faculty of Arts & Science, New York University, New York, NY 10003, USA
8 Department of Pharmaceutics, Wuya College of Innovation, Shenyang Pharmaceutical University, Shenyang 110016, China
9 Joint International Research Laboratory of Intelligent Drug Delivery Systems, Ministry of Education, Shenyang 110016, China
* Author to whom correspondence should be addressed
Received: 01 Jun 2025 Accepted: 10 Oct 2025 Available Online: 11 Oct 2025 Published: 27 Oct 2025
Endothelial cells (ECs) are essential for vascular network formation and tissue homeostasis, yet the fields of tissue engineering and vascularized organoid generation still rely heavily on human umbilical vein ECs (HUVECs), which are venous, allogeneic, and challenging to mature fully. Human pluripotent stem cells (hPSCs) offer an autologous, developmentally flexible alternative, but most differentiation protocols require fluorescence-activated cell sorting, limiting scalability. Here, we present a streamlined method that produces highly pure ECs directly from human embryonic stem cells (hESCs) without cell sorting. Extending Wnt pathway activation with CHIR99021 for three days maximizes mesoderm induction, while a brief Notch inhibition using DAPT during the specification phase suppresses smooth muscle cell commitment. The result is over 90% CD31+ CD144+ ECs that display classic cobblestone morphology, robust DiI-acetylated LDL uptake, and capillary-like sprouting comparable to HUVECs. Bulk RNA barcoding and sequencing segregates the hESC-derived ECs from both HUVECs and undifferentiated hESCs and uncovers an artery-enriched transcriptome: NOTCH1, DLL4, and CXCR4 are up-regulated, whereas venous markers (EPHB4, NRP2) are reduced. Enrichment of Notch-responsive pathways further supports an arterial-like identity. Although several adult functional genes (e.g., vWF, NOS3) are expressed at lower levels than in HUVECs, the protocol delivers a scalable source of developmentally relevant ECs ideal for vascularizing organoids derived from the same hPSCs and for future applications in drug screening, disease modeling, and cell-based vascular therapies.
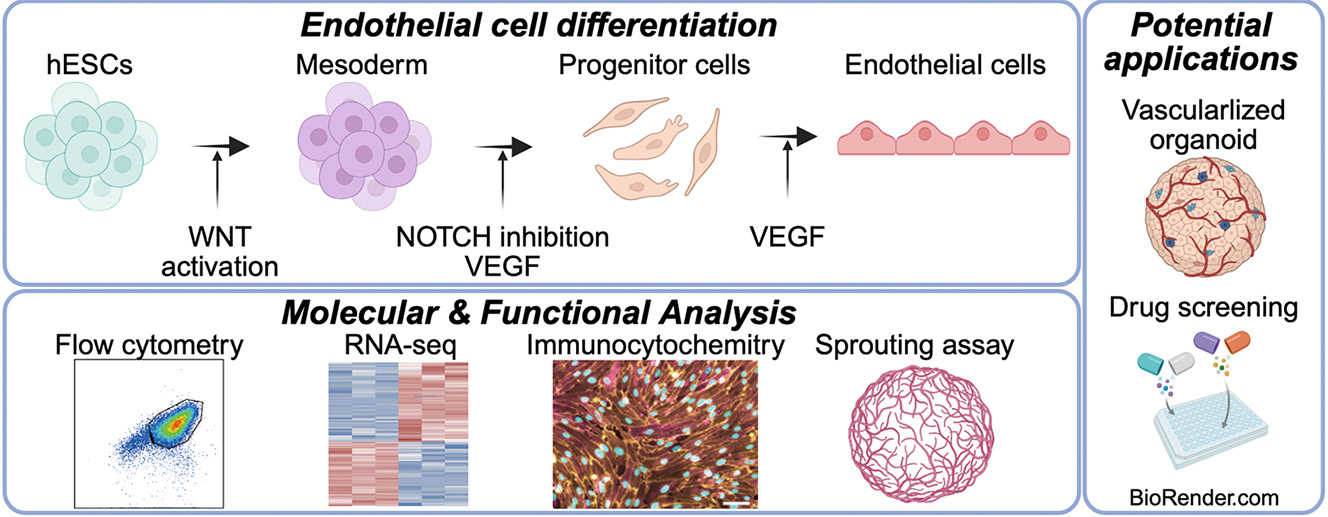
Endothelial cells (ECs) facilitate the differentiation and maturation of parenchymal cells in tissues through paracrine signaling and direct interactions during the early stages of organogenesis [1,2,3,4,5]. Subsequently, these cells line the entire vascular system, forming a dynamic interface critical for transporting oxygen, nutrients, and signaling molecules, as well as for immune surveillance and response, ensuring tissue viability and function [6]. Given these multifaceted and vital roles, the availability of functional human ECs is a cornerstone for advancing engineered human tissues, such as organoids that recapitulate in vivo complexities, and for developing novel strategies in regenerative medicine and drug discovery [7]. Human pluripotent stem cells (hPSCs), including human embryonic stem cells (hESCs) [8] and human induced pluripotent stem cells (hiPSCs) [9], have emerged as a promising and scalable source for generating a wide array of cell types, including those required for complex organoid assembly [7]. For vascularization strategies in engineered tissues, human umbilical vein endothelial cells (HUVECs) have historically been a common choice due to their relative ease of isolation and commercial availability. However, the utility of HUVECs in sophisticated tissue engineering and regenerative applications is increasingly recognized as limited. HUVECs often exhibit a limited capacity to support the differentiation and maturation of co-cultured parenchymal cells compared to primary human endothelial or organ-specific microvascular cells [7,10]. Their inherent venous identity further makes them suboptimal for constructing arterial or organ-specific vascular networks, which are critical for recapitulating physiological tissue function [11,12]. Moreover, the allogeneic nature of HUVECs poses substantial immunological challenges for engineered tissues intended for therapeutic transplantation, potentially resulting in graft rejection and limiting clinical translatability [13]. To address these limitations, ECs differentiated from patient-specific or HLA-matched hPSCs are gaining prominence as superior alternatives for organoid vascularization and regenerative therapies [14,15]. hPSC-ECs offer the potential for autologous or immunologically compatible cell sources, tunable differentiation towards specific endothelial subtypes (e.g., arterial, venous, lymphatic, or organ-specific), and robust scalability [4,5,16,17]. However, a persistent challenge in the field has been the inefficiency and heterogeneity of many current EC differentiation protocols. Many established methods yield EC populations (identified by CD31/PECAM1 and CD144/VE-Cadherin co-expression) with purities often below 80%, necessitating subsequent enrichment steps such as fluorescence-activated cell sorting (FACS) or magnetic-activated cell sorting (MACS) [16,18]. While effective, these purification procedures add substantial time, labor, and cost to the cell production process and may adversely affect cell viability and function, thereby limiting the scalability necessary for large-scale tissue engineering, drug screening, and clinical applications [19]. The establishment of a high-efficiency differentiation protocol that enables direct, sort-free induction of ECs from hPSCs is therefore of paramount importance, considering factors such as cost, time, and scalability. Early protocols explored the direct differentiation of hPSCs into ECs by activating key signaling pathways such as MEK/ERK and BMP4, achieving modest yields of CD31+ CD34+ cells (e.g., ~15%) [20]. Subsequent research underscored the pivotal role of precise temporal modulation of WNT signaling during mesoderm specification, revealing that transient WNT activation followed by its inhibition can significantly steer mesodermal progenitors towards an endothelial fate [16,21]. Furthermore, the interplay between VEGF signaling, essential for endothelial proliferation and survival, and Notch signaling has been identified as a critical regulatory axis. Specifically, Notch inhibition, often in conjunction with VEGF stimulation, has been reported to promote the specification and expansion of endothelial progenitors from hPSCs [18]. Despite these advancements, achieving direct differentiation efficiencies consistently exceeding 80–90% without sorting has remained elusive for many protocols, often due to incomplete lineage commitment or the emergence of off-target cell types [22,23,24]. Recent efforts continue to refine these strategies, focusing on optimized growth factor combinations, small molecule modulators, and defined culture conditions to improve purity and yield [19,23,25]. However, the need for robust, sort-free protocols that are easily adaptable and scalable remains paramount for widespread application. Here, we report a significantly refined and highly efficient protocol for the directed differentiation of hPSCs into ECs. By systematically optimizing the duration and timing of mesoderm induction using the selective GSK-3 inhibitor CHIR99021 and strategically introducing the Notch inhibitor DAPT during endothelial specification, we established an efficient differentiation protocol. This method consistently produces EC populations with over 90% purity for CD31 and CD144 co-expression, directly from hPSCs and without the need for cell sorting. We demonstrate that these hPSC-derived ECs exhibit characteristic endothelial morphology and robust sprouting angiogenesis in vitro, similar to HUVECs. They also possess a transcriptomic signature indicative of an arterial endothelial phenotype, characterized by enriched expression of arterial markers. This optimized, sort-free approach offers a scalable, cost-effective, and time-efficient means to generate large quantities of high-purity hPSC-ECs, providing a valuable and versatile cell source for constructing vascularized organoids, advancing regenerative medicine strategies, and developing more predictive drug discovery platforms.
2.1. Human Embryonic Stem Cell (hESC) Culture and Maintenance Human embryonic stem cells (hESCs), line H9 (WA09; RRID: CVCL_9773), were obtained from WiCell Research Institute (Madison, WI, USA). KhES1-OCT4-EGFP (K1) hESCs were obtained from Kyoto University. For routine culture, hESC-certified Matrigel (Corning, Inc., Corning, NY, USA) was diluted 1:75 (v/v) with DMEM/F12 medium (Merck KGaA, Darmstadt, Germany) and used to coat culture dishes. The Matrigel solution was incubated in the dishes for 24 h at 4 °C. Before cell seeding, excess Matrigel was aspirated, and the coated surface was washed with fresh DMEM/F12 medium. hESCs were maintained in mTeSR1 medium (Stem Cell Technologies, Vancouver, Canada) supplemented with 1% (v/v) penicillin/streptomycin (Fujifilm Wako, Osaka, Japan). The medium was changed daily. For passaging, cells were first washed with Dulbecco’s phosphate-buffered saline (D-PBS; devoid of calcium and magnesium; Thermo Fisher Scientific, Waltham, MA, USA). Dissociation was achieved by incubating the cells with TrypLE Express (Thermo Fisher Scientific) for 3 min at 37 °C. The detached cells were harvested and pelleted by centrifugation at 200× g for 3 min. Cell pellets were subsequently resuspended in mTeSR1 medium. To enhance cell survival post-dissociation, mTeSR1 medium supplemented with 10 µM Y27632 ROCK inhibitor (Fujifilm Wako) was used for the first 24 h after passaging. Thereafter, cells were cultured in mTeSR1 medium without ROCK inhibitor, with daily medium replacement. Cells were maintained for a maximum of 10 passages. 2.2. Differentiation of hESCs Towards Endothelial Cells To initiate endothelial differentiation, cultured hESCs were washed with D-PBS and detached using TrypLE Express for 3 min at 37 °C. The addition of basal medium neutralized the reaction, and the cell suspension was transferred to a 15-mL conical tube. Cells were pelleted by centrifugation at 200× g for 3 min, and the supernatant was discarded. The cell pellet was resuspended in mTeSR1 medium supplemented with 10 µM Y27632 and 1% (v/v) penicillin/streptomycin. Cells were seeded at a density of 5.3 × 104 cells cm−2 onto Matrigel-coated wells of Nunc Cell-Culture Treated Multidishes (Thermo Fisher Scientific). The cells were cultured in a humidified incubator at 37 °C with 5% CO2 for 24 h. Subsequently, the medium was replaced daily for 3 days with differentiation medium 1, consisting of DMEM/F12 (Merck KGaA) supplemented with 25 ng/mL human recombinant BMP4 (R&D Systems, Minneapolis, MN, USA), 8–16 µM CHIR99021 (concentration adjusted based on cell confluency and morphology; ReproCELL, Kanagawa, Japan), 50 ng mL−1 human recombinant bFGF (Fujifilm Wako), 1% (v/v) B27 supplement (Thermo Fisher Scientific), 1% (v/v) GlutaMax Supplement (Thermo Fisher Scientific), and 1% (v/v) penicillin/streptomycin. Following this, cells were cultured for an additional 3 days with daily medium changes using differentiation medium 2, composed of StemPro-34 SFM (Thermo Fisher Scientific) supplemented with 100 ng mL−1 human recombinant VEGF-A165 (Fujifilm Wako), 10 µM DAPT (AdipoGen Life Sciences, Inc., San Diego, CA, USA), 2 µM Forskolin (Tokyo Chemical Industry Co., Ltd., Tokyo, Japan), 1% (v/v) GlutaMax Supplement, and 1% (v/v) penicillin/streptomycin. Finally, cells were matured for 2–5 days in EGM Endothelial Cell Growth Medium BulletKit (Lonza, Basel, Switzerland) supplemented with 100 ng mL−1 VEGF-A165 and 1% (v/v) penicillin/streptomycin, with daily medium changes. If excessive cell proliferation was observed on day 10 of differentiation (corresponding to day 4 of EGM treatment), the medium change was omitted for that day. 2.3. Culture of Human Umbilical Vein Endothelial Cells (HUVECs) HUVECs (KAC Co., Ltd., Kyoto, Japan) were cultured in EGM Endothelial Cell Growth Medium BulletKit (Lonza) supplemented with 100 ng/mL VEGF-A165 and 1% (v/v) penicillin/streptomycin. For passaging, HUVECs were washed with D-PBS (devoid of calcium and magnesium) and dissociated using 0.02% EDTA/0.1% trypsin solution (Kohjin-bio, Saitama, Japan) for 3 min at 37 °C. Detachment was neutralized by the addition of EGM medium containing 1 mg mL−1 trypsin inhibitor from soybean (Fujifilm Wako) in PBS supplemented with 1% (v/v) penicillin/streptomycin. The cells were pelleted by centrifugation at 200× g for 3 min, resuspended, and seeded in fresh EGM medium. 2.4. Culture of Caco-2 Caco-2 human colorectal adenocarcinoma cell lines were obtained from the American Type Culture Collection (ATCC). Cells were maintained in DMEM-low glucose (Sigma-Aldrich, St. Louis, MO, USA) supplemented with 10% (v/v) fetal bovine serum (FBS, Cell Culture Bioscience), 1% (v/v) nonessential amino acids (Thermo Fisher Scientific), and 1% (v/v) penicillin/streptomycin at 37 °C with 5% (v/v) CO2. Cells were passaged using trypsin/EDTA (0.04%/0.03% [v/v]) solution every 3 and 5 days and at ratios of 1:5 and 1:10, respectively. 2.5. Fluorescence-Activated Cell Sorting (FACS) Cells intended for FACS analysis were rinsed twice with PBS and harvested using 0.1% trypsin/EDTA, followed by neutralization with 1 mg mL−1 trypsin inhibitor. After cell counting, cells were diluted to a final concentration of 1 × 107 cells mL−1 in staining buffer (FBS) (BD Pharmingen, Franklin Lakes, NJ, USA). For antibody staining, 2.5 µL of fluorescence-labeled antibody (specific antibodies listed in Supplementary Table S1) was added to 50 µL of the cell suspension, and the mixture was incubated on ice (approximately 4 °C) for 30–60 min. Corresponding isotype controls were used at the same concentration as the primary antibodies for negative controls. After incubation, excess antibodies were removed by washing the cells with staining buffer, followed by centrifugation at 300× g for 3 min. The stained cell suspensions were analyzed and sorted using a FACSAria II SORP cell sorter (BD Biosciences, Franklin Lakes, NJ, USA). Data were analyzed using FlowJo software (v9; FlowJo, LLC, Ashland, OR, USA). 2.6. Immunocytochemistry Cells cultured on appropriate surfaces were fixed with 4% paraformaldehyde (PFA) in D-PBS (devoid of calcium and magnesium) for 20 min at 25 °C. Following fixation, cells were permeabilized with 0.1% (v/v) Triton X-100 in D-PBS for 5 min at 25 °C. Non-specific binding sites were blocked by incubating the cells in blocking buffer [D-PBS containing 5% (v/v) normal goat serum (Maravai Life Sciences, San Diego, CA, USA), 5% (v/v) normal donkey serum (Jackson ImmunoResearch, West Grove, PA, USA), 3% (w/v) bovine serum albumin (BSA), essentially globulin-free (Merck KGaA), and 0.1% (v/v) Tween-20 (Nacalai Tesque, Kyoto, Japan)] at 4 °C for 16 h. Cells were then incubated with primary antibodies, diluted in blocking buffer as detailed in Supplementary Table S2, at 4 °C for 16 h. Subsequently, cells were washed and incubated with appropriate secondary antibodies, diluted as described in Supplementary Table S2, at 37 °C for 60 min. Nuclei were counterstained with DAPI (Fujifilm Wako) at 25 °C for 30 min. 2.7. Acetylated Low-Density Lipoprotein Uptake Assay Cells were washed once with D-PBS (devoid of calcium and magnesium) and then incubated with 10 μg mL−1 1,1′-dioctadecyl-3,3,3′,3′-tetramethylindocarbocyanine (Dil)-labeled acetylated low-density lipoprotein (Dil-AcLDL) (Thermo Fisher Scientific) diluted in endothelial basal medium (EBM) supplemented with 1% bovine serum albumin (BSA) at 37 °C in 5% CO2 for 3 h. After incubation, cells were washed twice with D-PBS at room temperature to remove unbound Dil-AcLDL. For flow cytometric analysis, cells were detached by incubation with 0.02% EDTA/0.1% trypsin solution at 37 °C for 3 min, collected in EBM medium, and immediately subjected to flow cytometry. For fluorescence microscopy, cells were fixed with 4% PFA for 15 min at 25 °C, followed by D-PBS washing. Nuclei were counterstained with DAPI for 5 min, rewashed with D-PBS, and imaged using a fluorescence microscope to visualize Dil-AcLDL uptake. 2.8. Bulk RNA Barcoding and Sequencing Total RNA was purified using the RNeasy Micro Kit (Qiagen, Hilden, Germany) and stored at −80 °C until library preparation. Bulk RNA barcoding and sequencing (BRB-seq) [26] was libraries were generated with the following modifications. First-strand cDNA was synthesized with an oligo-dT primer, and double-stranded cDNA was produced using the Second Strand Synthesis Module (New England Biolabs, Ipswich, MA, USA). Tagmentation was carried out with an in-house MEDS-B Tn5 transposase [27,28], followed by 10 PCR cycles using Phusion High-Fidelity DNA Polymerase (Thermo Fisher Scientific). Paired-end sequencing (Read 2 length: 81 bp) was performed on an Illumina NovaSeq 6000 (Illumina, Inc., San Diego, CA, USA). 2.9. Bulk RNA Barcoding and Sequencing (BRB-Seq) and Data Processing Barcodes were extracted with UMI-tools v1.1.1 using: “umi_tools extract -I read1.fastq --read2-in=read2.fastq\ --bc-pattern=NNNNNNNNNNCCCCCCCCC --read2-stdout” Adaptor trimming, low-quality base removal, and length filtering (<20 bp) were conducted with Trim Galore v0.6.7. High-quality reads were aligned to the human reference genome (GRCh38) with HISAT2 v2.2.1, and gene-level counts were generated with feature Counts v2.0.1. Differential expression analysis was performed in DESeq2 v1.34.0 and iDEP.96 [29], applying |log2(fold change)| > 1 and adjusted p < 0.05 as significance thresholds. 2.10. Comparative RNA-Seq Analysis GSEA (version 4.4.0) was used to compare the transcriptomic profile of our protocol-derived ECs to those of in vivo ECs [30]. Specifically, we used Macrovascular endothelial cells (Cluster 10) and Liver sinusoidal endothelial cells (Cluster 9) from the study. The signature gene sets for these clusters were defined by the reported marker genes [30], with a mean expression value (mean.cl) of ≥1. This resulted in gene sets of 91 and 90 genes for Cluster 10 and Cluster 9, respectively. For our protocol-derived ECs and HUVECs, we calculated the raw gene counts and then converted them to log2(Transcript per million + 1). For comparison, we used data from hiPSC-derived ECs [31] and ESC-derived ECs [32]. We obtained the Seurat object for Paik et al.’s data from NCBI GEO (accession number GSE116555), extracted the endothelial cells (cluster 4, resolution = 0.2) at day 12 post-differentiation, and performed pseudo-bulk processing of gene expression. We acquired the scRNA-seq raw data for McCracken et al.’s study from NCBI GEO (accession number GSE131736). Data analysis was performed using R (v4.4.3) and the Seurat package (v5.3.0), with Harmony (v1.2.3) applied for batch effect correction. After clustering with a resolution of 0.5, we isolated the ENPP2-positive cell population from the day 8 samples, as described in the McCracken et al. paper, and performed pseudo-bulk processing. To ensure a consistent GSEA analysis, we created a gene rank for each dataset using the expression profile of the ESCs from our study as a control. We used “Diff of Classes” as the GSEA ranking metric. 2.11. Endothelial Sprouting Assay Dishes were coated with 100 µL of Geltrex (Thermo Fisher Scientific) and incubated at 37 °C for 30 min. Cells were seeded in EGM™ Endothelial Cell Growth Medium BulletKit and cultured for 48 h. Sprouting morphology characteristic of endothelial cells was then assessed microscopically. 2.12. Fluorescence Imaging Samples were imaged on a Nikon ECLIPSE Ti inverted fluorescence microscope (Nikon Instruments, Tokyo, Japan) equipped with a CFI Plan Fluor 10×/0.30 NA objective, ORCA-R2 CCD camera (Hamamatsu Photonics, Shizuoka, Japan), Intensilight mercury lamp (Nikon), motorized XYZ stage (Ti-S-ER with encoders), and DAPI, GFP HYQ, and TRITC filter cubes (Nikon). 2.13. Statistical Analysis Statistical analyses were conducted in R v4.0.2. One-way ANOVA followed by Tukey’s post-hoc test or two-way ANOVA with Welch’s correction was used as appropriate. Differences were considered significant at p < 0.05.
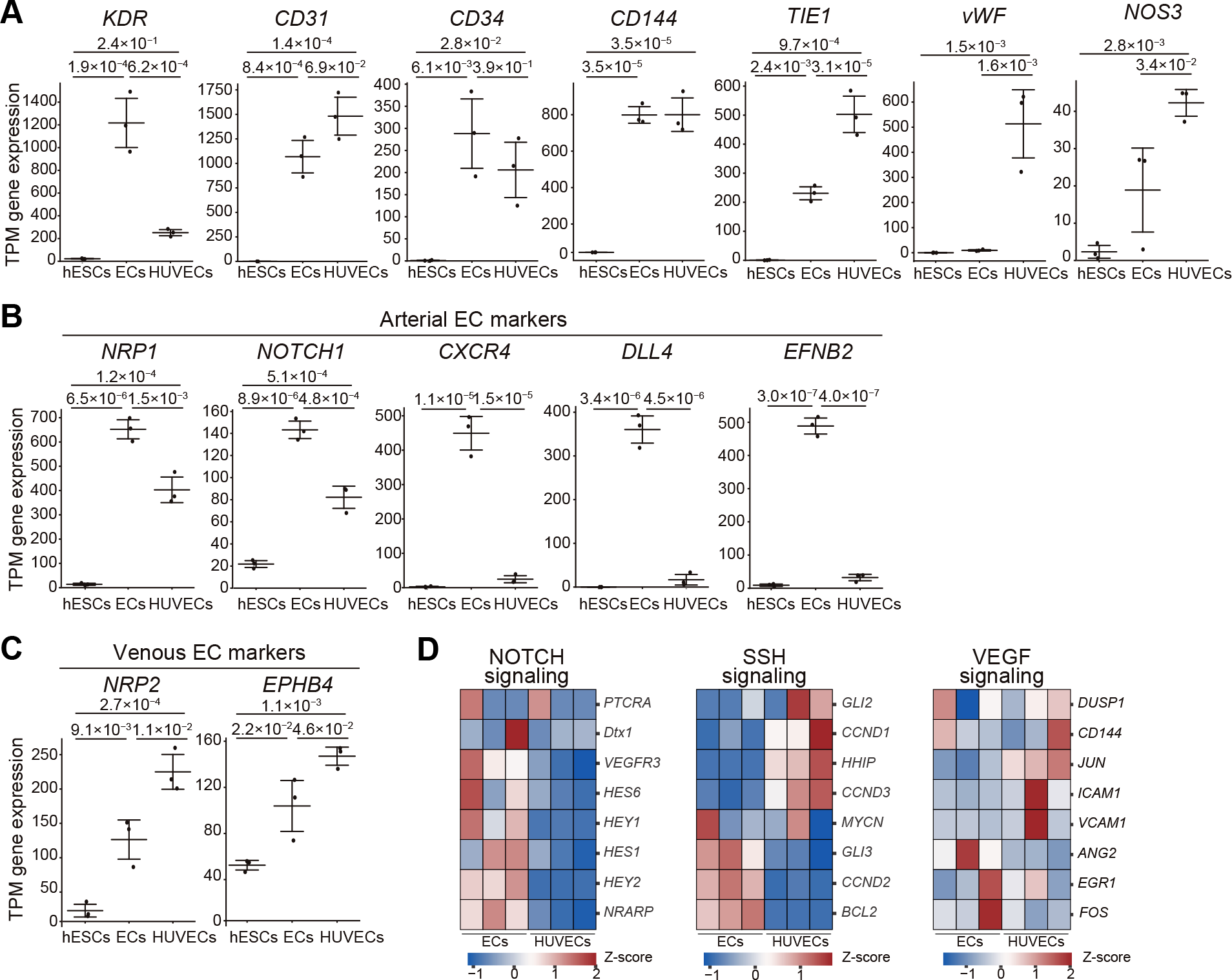
3.1. Optimization of hPSC Differentiation into Endothelial Cells Through Notch Inhibition and Increased Seeding Density To optimize endothelial differentiation from human pluripotent stem cells (hPSCs), we modified a previously published protocol by Takebe et al. [22] (Supplementary Figure S1A). Our refined approach involved optimizing the concentration of CHIR99021, a selective GSK-3 inhibitor for mesoderm induction, and DAPT, a γ-secretase inhibitor that promotes endothelial specification by inhibiting Notch signaling (Figure 1A). The fine-tuned protocol successfully generated ECs differentiated from H9 hESCs that displayed a typical cobblestone endothelial morphology (Figure 1B). In contrast, the original protocol yielded cells with more fibroblast-like morphology (Supplementary Figure S1B). We further investigated the impact of the initial cell seeding density on EC differentiation efficiency. While no noticeable morphological differences were observed across the tested densities (Supplementary Figure S1C), increasing the density from 1.9 × 104 to 5.3 × 104 cells cm−2 substantially improved the purity of the resulting EC population. Specifically, the percentage of cells co-expressing the endothelial markers CD144 and CD31 increased from 68.9% to 90.0% (Figure 1C and Supplementary Figure S1D). For comparison, undifferentiated H9 hESCs (negative control) contained only 0.40% CD144+ CD31+ cells, whereas HUVECs were 97.0% positive for these markers. Notably, our protocol consistently yielded a highly pure EC population without the need for subsequent cell sorting. This stands in stark contrast to the original method, which produced a mixed population containing only approximately 10% CD144+ CD31+ cells (Figure 1C). The robustness of our protocol was confirmed using a different hESC line, K1, which achieved a differentiation purity of 98.9% CD144+ CD31+ cells (Supplementary Figure S2). Collectively, these results demonstrate that our refined protocol provides a highly efficient and reliable method for directing hPSC differentiation into a pure population of endothelial cells. 3.2. hPSC-Derived Cells Express Endothelial Markers and Exhibit Angiogenic Potential To validate the endothelial identity of the cells generated from our protocol, we performed immunofluorescence staining (Figure 2A and Supplementary Figure S3A). The H9 hESC-derived ECs showed robust and uniform expression of the canonical endothelial markers CD31, CD144, vascular endothelial growth factor receptor 2 (VEGFR2), and von Willebrand Factor (vWF). This expression profile was comparable to that of our positive control, HUVECs, whereas the undifferentiated H9 hESCs (negative control) were negative for these markers. In contrast, only 0.023% of H9 hESC-derived ECs expressed the epithelial marker EpCAM, confirming a limited epithelial cell population after EC differentiation from hESCs (Supplementary Figure S3B). We next evaluated the functional properties of the hESC-derived ECs. The cells exhibited the characteristic endothelial function of DiI-acetylated low-density lipoprotein (DiI-AcLDL) uptake, as visualized by fluorescence microscopy (Figure 2B) and quantified by flow cytometry (Figure 2C). This capability was comparable to that of HUVECs and was absent in undifferentiated H9 hESCs. Furthermore, when tested for angiogenic potential in a three-dimensional (3D) sprouting assay, the hESC-derived ECs successfully formed complex, capillary-like sprouts, mimicking the behavior of HUVECs. As expected, mesenchymal control cells did not create these structures under the same conditions (Figure 2D). Taken together, these results demonstrate that our differentiation protocol yields cells that are not only phenotypically but also functionally characteristic of bona fide endothelial cells. 3.3. BRB-Seq Reveals Distinct Transcriptional Profiles in hPSC-Derived ECs and HUVECs BRB-seq followed by principal-component analysis cleanly separated H9 hESC-derived ECs, undifferentiated H9 hESCs, and HUVECs into three clusters (Figure 3A). PC1 (64% variance) reflected the pluripotent-to-endothelial transition, whereas PC2 (24% variance) distinguished H9 hESC-derived ECs from HUVECs. K-means clustering of the 1000 most variable genes (k = 4) identified four gene sets with distinct pathway enrichments (Figure 3B, Table 1, and Supplementary Tables S1 and S3), such as Cluster A (213 genes), enriched in HUVECs and highlighted adhesion/migration pathways (e.g., CLDN11, COL8A1, and LYVE1), Cluster B (320 genes), shared by both endothelial populations and contained classic EC genes (CD31, and CDH5) and “blood-vessel development” pathways, Cluster C (341 genes) marked pluripotent hESCs (SOX2, and POU5F1) with “cell-differentiation” signatures, and Cluster D (126 genes) specific to hPSC-derived ECs (COL1A1, and SOX6) and enriched for morphogenesis/developmental processes. K-means clustering (k = 4) of the 1000 most variable genes from BRB-seq transcriptional comparison of hESC-derived endothelial cells, hESCs, and HUVECs revealed four gene sets with distinct pathway enrichments. Thus, H9 hESC-derived ECs and HUVECs share a core endothelial program yet maintain distinct transcriptional identities. 3.4. Arterial Gene Programs Are Up-Regulated in hPSC-Derived ECs Differential-expression analysis between H9 hESC-derived ECs and HUVECs revealed higher expression of KDR and T-cadherin in the former (Figure 4A,B). Enrichment analysis of up-regulated genes pointed to “blood-vessel development” and “tube morphogenesis” pathways, featuring arterial markers such as NOTCH1, CXCR4, and DLL4. Conversely, HUVECs showed higher expression of genes linked to immune response and vascular homeostasis (e.g., VCAM1 and ACE) (Figure 4C; Supplementary Table S2). We confirmed that typical EC surface markers (KDR, CD31, CD34, and CD144) were similar between hPSC-derived ECs and HUVECs, but hPSC-derived ECs expressed lower levels of TIE1, vWF, and NOS3 (Figure 5A). Moreover, the expression of arterial-specific genes (NRP1, NOTCH1, CXCR4, DLL4, and EFNB2) was significantly higher in H9 hESC-derived ECs, whereas venous genes (NRP2 and EPHB4) were considerably lower (Figure 5B,C). Pathway-level inspection showed more vigorous Notch-signaling activity in H9 hESC-derived ECs, whereas Shh and VEGF-target gene sets did not differ markedly (Figure 5D). Collectively, these data indicate that our protocol drives hPSCs toward an arterial-like endothelial phenotype, distinct from the venous identity of HUVECs. 3.5. Transcriptomic Analysis Defines hESC-Derived ECs as a Macrovascular Subtype To benchmark the identity of our hESC-derived ECs against in vivo ECs, we performed a comparative transcriptomic analysis (Figure 6). We used Gene Set Enrichment Analysis (GSEA) to compare our cells with endothelial subtype signatures identified in a single-cell human liver atlas [30]. Our analysis revealed that the H9 hESC-derived ECs are closer to macrovascular endothelial cells (Figure 6A) than to liver sinusoidal endothelial cells (Figure 6B). When compared against undifferentiated hESCs, our ECs showed a more substantial enrichment for the macrovascular gene signature (Cluster 10) (Normalized Enrichment Score [NES] = 2.46, FDR = 0.000) than for the liver sinusoidal signature (Cluster 9) (NES = 2.25, FDR = 0.000). This expression profile was highly similar to that of HUVECs, which are of macrovascular origin and also showed a strong enrichment for the macrovascular gene set (ES = 0.77, NES = 2.64, FDR = 0.000). We further compared our hESC-derived ECs to two previously published datasets: ECs derived from human induced pluripotent stem cells (iPSCs) (Dataset 1; Paik et al., 2018) [31] and another from hESCs (Dataset 2; McCracken et al., 2020; Supplementary Material for cell barcodes) [32]. GSEA showed that the macrovascular gene signature was more enriched in both Dataset 1 (ES = 0.65, FDR = 0.000) and Dataset 2 (ES = 0.67, FDR = 0.000) relative to our hESC-derived ECs (Figure 6). Transcriptomic analysis confirms that our differentiation protocol yields endothelial cells with a distinct macrovascular identity, more closely resembling in vivo macrovascular cells than liver sinusoidal cells.![Figure 1: Efficient differentiation of hESCs into endothelial cells. (A) Schematic illustration of the protocol to generate endothelial cells (ECs) from human embryonic stem cells (hESCs): brief Wnt activation with CHIR99021 followed by notch inhibition with DAPT. (B) Phase-contrast micrographs of cells during differentiation from H9 hESCs to ECs. Scale bar, 100 µm. (C) Representative flow-cytometric graphs for CD144 (VE-cadherin) and CD (PECAM-1) in ECs. Shown left to right: H9 hESC-derived ESCs in this study, H9 hESC-derived ECs with the previous protocol [22], undifferentiated H9 hESCs as a negative control, and HUVECs as a positive control. Percentiles of CD144+ CD31+ cells are noted for each graph.](/uploads/source/articles/cell-therapy-engineering-connect/2025/volume1/20250006/image001.png)


Cluster
Pathways
adj.Pval
nGenes
Contained Top 5 Genes
A
Cell adhesion
1.23 × 10−8
46
CLDN11, IGFBP7, COL8A1, LYVE1, LGALS1
Cell migration
1.57 × 10−8
47
NR2F2. RAC2, NTN4, MTUS1, CCL2
Cell activation
2.21 × 10−7
41
PTX3, ANPEP, IL1RL1, RAC2, MFNG
B
Blood vessel development
3.26 × 10−44
85
PECAM1(CD31), ESM1, CDH5, THBS1, CLEC14A
Vasculature development
3.26 × 10−44
87
PECAM1(CD31), ESM1, CDH5, THBS1, CLEC14A
Blood vessel morphogenesis
9.31 × 10−41
77
ESM1, CDH5, THBS1, CLEC14A, CLDN5
C
Nervous system development
6.10 × 10−17
106
LIN28A, SOX2, CD24, ZIC2, SFRP2
Cell differentiation
7.21 × 10−11
125
POU5F1, LIN28A, SOX2, CD24, ZIC2
Neuron differentiation
7.21 × 10−11
66
LIN28A, SOX2, TUBB2B, SOX11, DNMT3B
D
Anatomical structure morphogenesis
1.00 × 10−10
52
CPM, COL1A1, COL11A1, DSP, SOX6
Cellular developmental process
6.72 × 10−7
57
DIO3, COL1A1, STMN2, COL11A1, DSP
Regulation of multicellular organismal process
1.41 × 10−6
43
COL1A1, DSP, SOX6, NPR3, EFNB2

![Figure 6: Transcriptomic analysis aligns hESC-derived ECs with a macrovascular endothelial cell identity. Gene Set Enrichment Analysis (GSEA) was used to compare H9 hESC-derived ECs, HUVECs, and two previously published EC datasets against endothelial gene signatures from a human liver cell atlas [30]. The comparison datasets are dataset 1 (iPSC-derived ECs [31]) and dataset 2 (ESC-derived ECs [32]). (A) GSEA plots showing the enrichment of the macrovascular endothelial cell gene signature (Cluster 10). The analysis reveals strong enrichment in hESC-derived ECs and HUVECs when compared to hESCs. (B) GSEA plots showing the enrichment of the liver sinusoidal endothelial cell gene signature (cluster 9). Enrichment scores for this gene set are lower across all comparisons than for the macrovascular gene set in (A), suggesting our hESC-derived ECs are more similar to a macrovascular gene expression signature—ES, Enrichment Score; NES, Normalized Enrichment Score; FDR, False Discovery Rate.](/uploads/source/articles/cell-therapy-engineering-connect/2025/volume1/20250006/image006.png)
In this study, we achieved highly efficient differentiation of hESCs into ECs with over 90% purity, using a protocol involving WNT pathway activation (CHIR99021) followed by NOTCH signaling inhibition (DAPT). As in vivo studies have shown, WNT signaling during mesoderm induction plays a critical role in EC differentiation [21], making the optimization of CHIR99021 treatment duration essential when differentiation efficiency is suboptimal. NOTCH signaling inhibitors promote differentiation into vascular endothelial progenitors by suppressing smooth muscle cell lineage commitment [20,21]. This refined method consistently yields a nearly pure population of ECs directly from culture, eliminating the need for subsequent cell sorting. This is a significant improvement over previous protocols, which often yielded heterogeneous cell mixtures with lower EC differentiation efficiencies, typically only ~50–70% of cells attaining an EC phenotype without purification [32,33]. For example, single-cell analyses of conventional 8-day directed differentiation showed only ~66% of cells co-expressing EC markers, with the remainder adopting non-endothelial fates [32]. Our approach overcomes this limitation, producing more than 90% ECs in one step, whereas earlier methods generally required cell sorting to enrich the endothelial fraction. Functionally, the resulting hESC-ECs display the classic cobblestone morphology, efficiently take up Dil-AcLDL, and form capillary-like sprouts, providing clear evidence of endothelial identity in vitro [34]. To define their gene expression profile, bulk RNA barcoding and sequencing further revealed an artery-enriched transcriptional profile: NOTCH1, DLL4, CXCR4, and EFNB2 were elevated, whereas venous markers (EPHB4, and NRP2) were reduced. Notably, others have observed a similar arterial bias in hPSC-derived endothelium—for instance, Paik et al. identified an iPSC-EC subpopulation marked by GJA5 (Cx40), indicative of an arterial-like identity. [33]. This default arterial programming is standard in many PSC-EC protocols [19], likely because strong WNT activation coupled with abundant endogenous VEGF signaling drives arterial specification, overriding the brief NOTCH inhibition in our system. In line with this, our cells did not prominently express venous-fate genes, reflecting a tendency for hPSC-ECs to adopt arterial-like characteristics unless specific venous-inducing cues are provided [17]. To further benchmark our hESC-derived ECs against their in vivo counterparts, we performed GSEA using a single-cell human liver atlas [30]. This comparative analysis revealed that the gene signature of our hESC-derived EC is significantly more enriched for macrovascular endothelial cells than for liver sinusoidal endothelial cells. In other words, the transcriptome of our hESC-ECs more closely resembles large-vessel endothelium rather than organ-specialized microvasculature, which is consistent with their artery-enriched signature and robust functional traits. This finding aligns with reports that PSC-derived EC products have a molecular identity distinct from organ-specific endothelium [32]. For instance, a recent single-cell study noted that the global transcriptional architecture of hESC-ECs differs from that of freshly isolated tissue-specific ECs [32,35,36,37]. Our results confirm that our protocol generates ECs with a distinct macrovascular identity, matching their high expression of arterial markers and firm performance in endothelial functional assays. The high yield and arterial bias of these ECs make them highly suitable for constructing vascularized organoids for disease modeling and drug discovery. In such systems, the presence of perfusable microvessels enhances organoid size, viability, and physiological relevance by facilitating nutrient delivery and waste removal through blood vessel–like networks [10,23,38]. Indeed, large-scale organoid studies have demonstrated that incorporating iPSC-derived endothelial cells can significantly enhance tissue maturation and function—for example, Takebe et al. generated vascularized liver buds entirely from human iPSCs and observed improved hepatic functions and engraftment in vivo [22]. The protocol also supports autologous cell therapy applications, because our ECs can be generated in large numbers from pluripotent sources, including patient-specific iPSCs. Patient-derived induced pluripotent stem cell–derived endothelial cells (iPSC-ECs) have demonstrated significant therapeutic potential in various ischemic vascular disease models. These include enhanced perfusion and vascular repair in hindlimb ischemia [39], attenuation of white matter ischemic injury and promotion of remyelination following cerebral infarction [40], and restoration of choroidal blood flow with recovery of visual function in ocular ischemic injury models [41]. Because the cells can be generated in large numbers from pluripotent sources, including patient-specific iPSCs, the protocol also supports autologous cell therapies and the endothelialization of engineered grafts for ischemic disease or tissue repair [42]. By providing a ready supply of high-purity ECs with a stable arterial phenotype, our method could thus accelerate the development of personalized vascular grafts and improve the vascular integration of tissue-engineered implants. However, some limitations remain. First, the cells remain functionally immature, likely because they develop in static 2D culture lacking hemodynamic shear, extracellular matrix architecture, and crosstalk with mural or blood cells. It is well known that biomechanical forces play a key role in endothelial maturation—for example, exposing iPSC-derived ECs to laminar shear stress in a flow bioreactor rapidly induces an in vivo-like arterial phenotype with enhanced anti-thrombotic and anti-inflammatory gene expression [43,44]. Refinements will be needed to derive venous endothelial cells or more uniformly arterial cells as required for specific applications. Finally, we have not yet assessed long-term stability, tumorigenic risk, or in vivo engraftment performance of these hESC-ECs. Such evaluations will require further studies using chemically defined, clinically compliant reagents to ensure safety and functionality for therapeutic use. Future work should focus on generating fully mature arterial or venous endothelial subtypes, potentially through stage-specific modulation of Notch and VEGF signaling pathways, exposure to physiological levels of laminar shear stress, or co-culture with supporting cells (such as pericytes or smooth muscle cells) to provide inductive signals. For instance, precisely timed activation or inhibition of Notch signaling during differentiation could direct cells toward arterial or venous fates. At the same time, post-differentiation conditioning under flow or within 3D matrices is expected to drive further maturation. Extended culture under laminar flow conditions can markedly boost the expression of arterial-enriched genes (like EFNB2, DLL4, NOS3) and enhance barrier integrity and anti-thrombotic function of PSC-ECs. Likewise, embedding the cells in a 3D ECM or co-culturing with stromal cells may provide biochemical and mechanical cues that upregulate vWF, tighten junctions, and improve alignment and response to inflammatory stimuli, making the cells more functionally equivalent to their in vivo counterparts. Refining these parameters is expected to yield endothelial cells that are not only abundant and phenotypically well-defined but also highly functional and mature, thereby significantly facilitating downstream applications. The ability to efficiently leverage combinations of biochemical and biophysical cues to fine-tune EC fate and maturity will significantly accelerate the development of vascularized organoids, reliable drug-screening platforms, and next-generation vascular therapies. By achieving closer parity with primary human endothelium, PSC-derived ECs produced via improved protocols like ours can better realize their promise in regenerative medicine and tissue engineering.
We established an efficient method: brief WNT activation with CHIR99021, followed by Notch inhibition with DAPT, to differentiate hESCs into endothelial cells, resulting in over 90% purity without cell sorting. The cells show key endothelial behaviors (cobblestone shape, Dil-AcLDL uptake, sprouting) and an arterial-leaning gene profile. Although some mature markers remain lower than in primary HUVECs, the protocol yields large numbers of functional ECs suitable for vascularized organoids, drug testing, and future cell-therapy studies. Fine-tuning NOTCH/VEGF signaling and adding flow or co-culture steps should further mature the cells and allow selective arterial or venous production.
| ACE | Angiotensin I Converting Enzyme |
| ANOVA | Analysis of Variance |
| BRB-seq | Bulk RNA Barcoding and Sequencing |
| CD | Cluster of Differentiation |
| CDH5 | Cadherin 5 (VE-cadherin) |
| CLDN11 | Claudin 11 |
| COL8A1 | Collagen Type VIII Alpha 1 Chain |
| CXCR4 | C-X-C Motif Chemokine Receptor 4 |
| DLL4 | Delta Like Canonical Notch Ligand 4 |
| ECs | Endothelial Cells |
| ENPP2 | Ectonucleotide Pyrophosphatase/Phosphodiesterase 2 (Autotaxin) |
| EpCAM | Epithelial Cell Adhesion Molecule |
| EPHB4 | EPH Receptor B4 |
| ES | Enrichment Score |
| FACS | Fluorescence-Activated Cell Sorting |
| FDR | False Discovery Rate |
| GJA5 | Gap Junction Protein Alpha 5 (Connexin 40) |
| GSEA | Gene Set Enrichment Analysis |
| hESCs | Human Embryonic Stem Cells |
| hiPSCs | Human Induced Pluripotent Stem Cells |
| hPSCs | Human Pluripotent Stem Cells |
| HUVECs | Human Umbilical Vein Endothelial Cells |
| KDR | Kinase Insert Domain Receptor (VEGFR2) |
| LYVE1 | Lymphatic Vessel Endothelial Hyaluronan Receptor 1 |
| MACS | Magnetic-Activated Cell Sorting |
| NES | Normalized Enrichment Score |
| NOS3 | Nitric Oxide Synthase 3 (eNOS) |
| NRP2 | Neuropilin 2 |
| SOX6 | SRY-Box Transcription Factor 6 |
| TIE1 | Tyro-sine Kinase with Immunoglobulin and EGF Like Do-mains 1 |
| VCAM1 | Vascular Cell Adhesion Molecule 1 |
| VEGFR2 | Vascular Endothelial Growth Factor Receptor 2 |
| vWF | von Willebrand Factor |
Conceptualization: K.Y., K.-i.K.; Methodology: K.Y., K.I., S.T., K.-i.K.; Investigation: K.Y., R.O., K.I., K.-i.K.; Visualization: K.Y., R.O., K.I.; Funding acquisition: K.Y., K.-i.K.; Project administration: K.-i.K.; Supervision: K.-i.K.; Writing—original draft, review & editing: K.Y., K.I., K.-i.K. All authors have read and agreed to the published version of the manuscript.
All BRB-seq data are available at the Gene Expression Omnibus (GSE298817).
All hESC experiments were conducted in accordance with the guidelines of the Ethics Committee of Kyoto University, [Kyoto, Japan] (Approval No. ES3-9, approved on 17 May 2010). HUVECs were obtained from KAC Co., Ltd. [Kyoto, Japan]. Caco-2 cells were obtained from the American Type Culture Collection (ATCC) [Manassas, VA, USA].
No consent for publication is required, as the manuscript does not involve any individual personal data, images, videos, or other materials that would necessitate consent. The research was conducted using established human embryonic stem cell lines (H9 and K1) available to the public.
K.-i.K. is a founder and CTO of Z24 Holdings. The other authors declare no competing financial or personal interests that could have influenced the work reported in this paper.
The Japan Society for the Promotion of Science (JSPS; 21H01728, 23H00260, and 24H00797 to K.-i.K., and 22KJ1987 to K.Y.) provided funding for this research. The Nakatani Foundation also provided financial support to K.Y.
The research was conducted in accordance with the principles of the Declaration of Helsinki. The use of human embryonic stem cells in this study was approved by the Ethics Committee of Kyoto University (Approval No. ES3-9).
We thank our lab members for their assistance. We also thank the iCeMS Analysis Center for access to the advanced microscopy and analytical instruments. ChatGPT o3 and Gemini 2.5 were used to improve English grammar and readability.
[1] Matsumoto, K.; Yoshitomi, H.; Rossant, J.; Zaret, K.S. Liver Organogenesis Promoted by Endothelial Cells Prior to Vascular Function. Science 2001, 294, 559–563. [CrossRef]
[2] Camp, J.G.; Sekine, K.; Gerber, T.; Loeffler-Wirth, H.; Binder, H.; Gac, M.; Kanton, S.; Kageyama, J.; Damm, G.; Seehofer, D.; et al. Multilineage Communication Regulates Human Liver Bud Development from Pluripotency. Nature 2017, 546, 533–538. [CrossRef]
[3] Carmeliet, P.; Jain, R.K. Angiogenesis in Cancer and Other Diseases. Nature 2000, 407, 249–257. [CrossRef]
[4] Pan, Z.; Yao, Q.; Kong, W.; Ma, X.; Tian, L.; Zhao, Y.; Zhu, S.; Chen, S.; Sun, M.; Liu, J.; et al. Generation of iPSC-Derived Human Venous Endothelial Cells for the Modeling of Vascular Malformations and Drug Discovery. Cell Stem Cell 2025, 32, 227–245.e9. [CrossRef] [PubMed]
[5] Ang, L.T.; Nguyen, A.T.; Liu, K.J.; Chen, A.; Xiong, X.; Curtis, M.; Martin, R.M.; Raftry, B.C.; Ng, C.Y.; Vogel, U.; et al. Generating Human Artery and Vein Cells From Pluripotent Stem Cells Highlights the Arterial Tropism of Nipah and Hendra Viruses. Cell 2022, 185, 2523–2541.e30. [CrossRef] [PubMed]
[6] Augustin, H.G.; Koh, G.Y. Organotypic Vasculature: From Descriptive Heterogeneity to Functional Pathophysiology. Science 2017, 357, eaal2379. [CrossRef]
[7] Takebe, T.; Sekine, K.; Enomura, M.; Koike, H.; Kimura, M.; Ogaeri, T.; Zhang, R.R.; Ueno, Y.; Zheng, Y.W.; Koike, N.; et al. Vascularized and Functional Human Liver from an iPSC-Derived Organ Bud Transplant. Nature 2013, 499, 481–484. [CrossRef]
[8] Thomson, J.A.; Itskovitz-Eldor, J.; Shapiro, S.S.; Waknitz, M.A.; Swiergiel, J.J.; Marshall, V.S.; Jones, J.M. Embryonic Stem Cell Lines Derived from Human Blastocysts. Science 1998, 282, 1145–1147. [CrossRef] [PubMed]
[9] Takahashi, K.; Tanabe, K.; Ohnuki, M.; Narita, M.; Ichisaka, T.; Tomoda, K.; Yamanaka, S. Induction of Pluripotent Stem Cells from Adult Human Fibroblasts by Defined Factors. Cell 2007, 131, 861–872. [CrossRef]
[10] Yap, K.K.; Gerrand, Y.W.; Dingle, A.M.; Yeoh, G.C.; Morrison, W.A.; Mitchell, G.M. Liver Sinusoidal Endothelial Cells Promote the Differentiation and Survival of Mouse Vascularised Hepatobiliary Organoids. Biomaterials 2020, 251, 120091. [CrossRef]
[11] Adams, R.H.; Alitalo, K. Molecular Regulation of Angiogenesis and Lymphangiogenesis. Nat. Rev. Mol. Cell Biol. 2007, 8, 464–478. [CrossRef]
[12] Potente, M.; Carmeliet, P. The Link Between Angiogenesis and Endothelial Metabolism. Annu. Rev. Physiol. 2017, 79, 43–66. [CrossRef]
[13] Scheiner, Z.S.; Talib, S.; Feigal, E.G. The Potential for Immunogenicity of Autologous Induced Pluripotent Stem Cell-Derived Therapies. J. Biol. Chem. 2014, 289, 4571–4577. [CrossRef]
[14] Macklin, B.L.; Gerecht, S. Bridging the Gap: Induced Pluripotent Stem Cell Derived Endothelial Cells for 3D Vascular Assembly. Curr. Opin. Chem. Eng. 2017, 15, 102–109. [CrossRef]
[15] Kirkeby, A.; Main, H.; Carpenter, M. Pluripotent Stem-Cell-Derived Therapies in Clinical Trial: A 2025 Update. Cell Stem Cell 2025, 32, 10–37. [CrossRef]
[16] Patsch, C.; Challet-Meylan, L.; Thoma, E.C.; Urich, E.; Heckel, T.; O’Sullivan, J.F.; Grainger, S.J.; Kapp, F.G.; Sun, L.; Christensen, K.; et al. Generation of Vascular Endothelial and Smooth Muscle Cells from Human Pluripotent Stem Cells. Nat. Cell Biol. 2015, 17, 994–1003. [CrossRef]
[17] Arora, S.; Yim, E.K.F.; Toh, Y.C. Environmental Specification of Pluripotent Stem Cell Derived Endothelial Cells Toward Arterial and Venous Subtypes. Front. Bioeng. Biotechnol. 2019, 7, 143. [CrossRef] [PubMed]
[18] Sahara, M.; Hansson, E.M.; Wernet, O.; Lui, K.O.; Später, D.; Chien, K.R. Manipulation of a VEGF-Notch Signaling Circuit Drives Formation of Functional Vascular Endothelial Progenitors from Human Pluripotent Stem Cells. Cell Res. 2014, 24, 820–841. [CrossRef] [PubMed]
[19] Farkas, S.; Simara, P.; Rehakova, D.; Veverkova, L.; Koutna, I. Endothelial Progenitor Cells Produced From Human Pluripotent Stem Cells by a Synergistic Combination of Cytokines, Small Compounds, and Serum-Free Medium. Front. Cell Dev. Biol. 2020, 8, 309. [CrossRef] [PubMed]
[20] Park, S.W.; Koh, Y.J.; Jeon, J.; Cho, Y.H.; Jang, M.J.; Kang, Y.; Kim, M.J.; Choi, C.; Cho, Y.S.; Chung, H.M.; et al. Efficient Differentiation of Human Pluripotent Stem Cells into Functional CD34+ Progenitor Cells by Combined Modulation of the MEK/ERK and BMP4 Signaling Pathways. Blood 2010, 116, 5762–5772. [CrossRef]
[21] Martin, B.L.; Kimelman, D. Canonical Wnt Signaling Dynamically Controls Multiple Stem Cell Fate Decisions during Vertebrate Body Formation. Dev. Cell 2012, 22, 223–232. [CrossRef] [PubMed]
[22] Takebe, T.; Sekine, K.; Kimura, M.; Yoshizawa, E.; Ayano, S.; Koido, M.; Funayama, S.; Nakanishi, N.; Hisai, T.; Kobayashi, T.; et al. Massive and Reproducible Production of Liver Buds Entirely from Human Pluripotent Stem Cells. Cell Rep. 2017, 21, 2661–2670. [CrossRef]
[23] Yang, L.; Han, Y.; Zhang, T.; Dong, X.; Ge, J.; Roy, A.; Zhu, J.; Lu, T.; Jeya Vandana, J.; de Silva, N.; et al. Human Vascularized Macrophage-Islet Organoids to Model Immune-Mediated Pancreatic β Cell Pyroptosis upon Viral Infection. Cell Stem Cell 2024, 31, 1612–1629.e8. [CrossRef]
[24] Orlova, V.V.; Van Den Hil, F.E.; Petrus-Reurer, S.; Drabsch, Y.; Ten Dijke, P.; Mummery, C.L. Generation, Expansion and Functional Analysis of Endothelial Cells and Pericytes Derived from Human Pluripotent Stem Cells. Nat. Protoc. 2014, 9, 1514–1531. [CrossRef]
[25] Batta, K.; Menegatti, S.; Garcia-Alegria, E.; Florkowska, M.; Lacaud, G.; Kouskoff, V. Concise Review: Recent Advances in the In Vitro Derivation of Blood Cell Populations. Stem Cells Transl. Med. 2016, 5, 1330–1337. [CrossRef]
[26] Alpern, D.; Gardeux, V.; Russeil, J.; Mangeat, B.; Meireles-Filho, A.C.A.; Breysse, R.; Hacker, D.; Deplancke, B. BRB-seq: Ultra-affordable High-throughput Transcriptomics Enabled by Bulk RNA Barcoding and Sequencing. Genome Biol. 2019, 20, 71. [CrossRef]
[27] Picelli, S.; Björklund, Å.K.; Reinius, B.; Sagasser, S.; Winberg, G.; Sandberg, R. Tn5 Transposase and Tagmentation Procedures for Massively Scaled Sequencing Projects. Genome Res. 2014, 24, 2033–2040. [CrossRef]
[28] Sato, S.; Arimura, Y.; Kujirai, T.; Harada, A.; Maehara, K.; Nogami, J.; Ohkawa, Y.; Kurumizaka, H. Biochemical Analysis of Nucleosome Targeting by Tn5 Transposase. Open Biol. 2019, 9, 190116. [CrossRef] [PubMed]
[29] Ge, S.X.; Son, E.W.; Yao, R. iDEP: An Integrated Web Application for Differential Expression and Pathway Analysis of RNA-Seq Data. BMC Bioinformatics 2018, 19, 1–24. [CrossRef]
[30] Aizarani, N.; Saviano, A.; Sagar; Mailly, L.; Durand, S.; Herman, J.S.; Pessaux, P.; Baumert, T.F.; Grün, D. A Human Liver Cell Atlas Reveals Heterogeneity and Epithelial Progenitors. Nature 2019, 572, 199–204. [CrossRef] [PubMed]
[31] Paik, D.T.; Tian, L.; Lee, J.; Sayed, N.; Chen, I.Y.; Rhee, S.; Rhee, J.W.; Kim, Y.; Wirka, R.C.; Buikema, J.W.; et al. Large-Scale Single-Cell RNA-Seq Reveals Molecular Signatures of Heterogeneous Populations of Human Induced Pluripotent Stem Cell-Derived Endothelial Cells. Circ. Res. 2018, 123, 443–450. [CrossRef]
[32] McCracken, I.R.; Taylor, R.S.; Kok, F.O.; de la Cuesta, F.; Dobie, R.; Henderson, B.E.P.; Mountford, J.C.; Caudrillier, A.; Henderson, N.C.; Ponting, C.P.; et al. Transcriptional Dynamics of Pluripotent Stem Cell-Derived Endothelial Cell Differentiation Revealed by Single-Cell RNA Sequencing. Eur. Heart J. 2020, 41, 1024–1036. [CrossRef] [PubMed]
[33] Yu, L.; Logsdon, D.; Pinzon-Arteaga, C.A.; Duan, J.; Ezashi, T.; Wei, Y.; Ribeiro Orsi, A.E.; Oura, S.; Liu, L.; Wang, L.; et al. Large-scale Production of Human Blastoids Amenable to Modeling Blastocyst Development and Maternal-Fetal Cross Talk. Cell Stem Cell 2023, 30, 1246–1261.e9. [CrossRef]
[34] Gu, M. Efficient Differentiation of Human Pluripotent Stem Cells to Endothelial Cells. Curr. Protoc. Hum. Genet. 2018, 98, e64. [CrossRef] [PubMed]
[35] Nguyen, J.; Lin, Y.Y.; Gerecht, S. The Next Generation of Endothelial Differentiation: Tissue-Specific ECs. Cell Stem Cell 2021, 28, 1188–1204. [CrossRef]
[36] Nolan, D.J.; Ginsberg, M.; Israely, E.; Palikuqi, B.; Poulos, M.G.; James, D.; Ding, B.S.; Schachterle, W.; Liu, Y.; Rosenwaks, Z.; et al. Molecular Signatures of Tissue-Specific Microvascular Endothelial Cell Heterogeneity in Organ Maintenance and Regeneration. Dev. Cell 2013, 26, 204–219. [CrossRef]
[37] Majid, Q.A.; Ghimire, B.R.; Merkely, B.; Randi, A.M.; Harding, S.E.; Talman, V.; Földes, G. Generation and Characterisation of Scalable and Stable Human Pluripotent Stem Cell-Derived Microvascular-Like Endothelial Cells for Cardiac Applications. Angiogenesis 2024, 27, 561–582. [CrossRef]
[38] Wimmer, R.A.; Leopoldi, A.; Aichinger, M.; Wick, N.; Hantusch, B.; Novatchkova, M.; Taubenschmid, J.; Hämmerle, M.; Esk, C.; Bagley, J.A.; et al. Human Blood Vessel Organoids as a Model of Diabetic Vasculopathy. Nature 2019, 565, 505–510. [CrossRef] [PubMed]
[39] Takii, A.; Tanabe, Y.; Li, W.; Shiomi, H.; Inoue, A.; Muramatsu, F.; Jia, W.; Takakura, N. CD157+ Vascular Endothelial Cells Derived from Human-Induced Pluripotent Stem Cells Have High Angiogenic Potential. Inflamm. Regen. 2025, 45, 14. [CrossRef]
[40] Xu, B.; Kurachi, M.; Shimauchi-Ohtaki, H.; Yoshimoto, Y.; Ishizaki, Y. Transplantation of iPS-Derived Vascular Endothelial Cells Improves White Matter Ischemic Damage. J. Neurochem. 2020, 153, 759–771. [CrossRef]
[41] Li, M.; Wang, P.; Huo, S.T.; Qiu, H.; Li, C.; Lin, S.; Guo, L.; Ji, Y.; Zhu, Y.; Liu, J.; et al. Human Pluripotent Stem Cells Derived Endothelial Cells Repair Choroidal Ischemia. Adv. Sci. 2024, 11, 2302940. [CrossRef]
[42] Rufaihah, A.J.; Huang, N.F.; Jamé, S.; Lee, J.C.; Nguyen, H.N.; Byers, B.; De, A.; Okogbaa, J.; Rollins, M.; Reijo-Pera, R.; et al. Endothelial Cells Derived From Human iPSCS Increase Capillary Density and Improve Perfusion in a Mouse Model of Peripheral Arterial Disease. Arterioscler. Thromb. Vasc. Biol. 2011, 31, 11. [CrossRef] [PubMed]
[43] Sivarapatna, A.; Ghaedi, M.; Le, A.V.; Mendez, J.J.; Qyang, Y.; Niklason, L.E. Arterial Specification of Endothelial Cells Derived from Human Induced Pluripotent Stem Cells in a Biomimetic Flow Bioreactor. Biomaterials 2015, 53, 621–633. [CrossRef] [PubMed]
[44] Kennedy, C.C.; Brown, E.E.; Abutaleb, N.O.; Truskey, G.A. Development and Application of Endothelial Cells Derived From Pluripotent Stem Cells in Microphysiological Systems Models. Front. Cardiovasc. Med. 2021, 8, 625016. [CrossRef] [PubMed]
Disclaimer/Publisher’s Note: The views expressed in this article are those of the author(s) and do not necessarily reflect the views of the publisher or editors. The publisher and editors assume no responsibility for any injury or damage resulting from the use of information contained herein.
We use cookies to improve your experience on our site. By continuing to use our site, you accept our use of cookies. Learn more